- Anserine ELISA Kits
- Avian ELISA Kits
- Bovine ELISA Kits
- Canine ELISA Kits
- Camel ELISA Kits
- Chicken ELISA Kits
- Monkey ELISA Kits
- Duck ELISA Kits
- Equine ELISA Kits
- Fish ELISA Kits
- Feline ELISA Kits
- Guinea Pig ELISA Kits
- Goose ELISA Kits
- Goat ELISA Kits
- Gymnuromys ELISA Kits
- Human ELISA Kits
- Hamster ELISA Kits
- Horse ELISA Kits
- Lizard ELISA Kits
- Mouse ELISA Kits
- Porcine ELISA Kits
- Pigeon ELISA Kits
- Rat ELISA Kits
- Sheep ELISA Kits
- Zebra ELISA Kits
- Deer ELISA Kits
- SPECIFICATION
- BACKGROUND
- DOCUMENTS
- REVIEW
ADDITIONAL INFORMATION
SYNONYMS
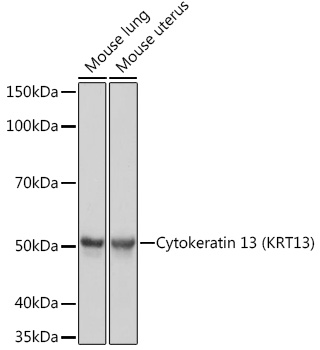
K13; CK13; WSN2; Cytokeratin 13 (KRT13)
REACTIVITY
Human,Mouse,Rat
TESTED APPLICATIONS
ELISA,WB,IHC-P,IF/ICC
RECOMMENDED DILUTION
WB,1:500 - 1:2000|IHC-P,1:50 - 1:200|IF/ICC,1:50 - 1:200
CALCULATED MW
50kDa
OBSERVED MW
50kDa
IMMUNOGEN
A synthetic peptide corresponding to a sequence within amino acids 100-200 of human Cytokeratin 13 (KRT13) (P13646).
POSITIVE SAMPLES
Mouse lung,Mouse uterus
CELLULAR LOCALIZATION
cytoskeleton,cytosol,extracellular exosome,intermediate filament cytoskeleton,nucleus,keratin filament
STORAGE BUFFER
Store at -20℃. Avoid freeze / thaw cycles.|Buffer: PBS with 0.02% sodium azide,0.05% BSA,50% glycerol,pH7.3.
BACKGROUND
The protein encoded by this gene is a member of the keratin gene family. The keratins are intermediate filament proteins responsible for the structural integrity of epithelial cells and are subdivided into cytokeratins and hair keratins. Most of the type I cytokeratins consist of acidic proteins which are arranged in pairs of heterotypic keratin chains. This type I cytokeratin is paired with keratin 4 and expressed in the suprabasal layers of non-cornified stratified epithelia. Mutations in this gene and keratin 4 have been associated with the autosomal dominant disorder White Sponge Nevus. The type I cytokeratins are clustered in a region of chromosome 17q21.2. Alternative splicing of this gene results in multiple transcript variants; however, not all variants have been described.
REVIEW
Neo Scientific welcomes feedback from its customers.
If you have used an our product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information.
If you have any additional inquiries please email technical services at info@neobiolab.com
Thank you for your support.